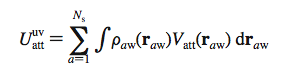Ammm:Molecular surfaces: Difference between revisions
Created page with "= Introduction = Calculating molecular surfaces is important and useful for structure-based drug design, representing molecules and computing inter-molecular interactions. It is also useful for displaying certain molecular properties on the surface, using color-coded and texture maps. Examples of such properties include atomic charge, electrostatic potential, hydrophobicity, polarizability, etc. = Types of Molecular Surfaces = File:Surfaces.png == Van der Waals S..." |
No edit summary |
||
| Line 350: | Line 350: | ||
= Further Reading = | = Further Reading = | ||
[[File:Lee-richards71.pdf]] | [[File:Lee-richards71.pdf | thumb]] | ||
[[File:Richards77.pdf]] | [[File:Richards77.pdf| thumb]] | ||
[[File:Connolly.pdf]] | [[File:Connolly.pdf| thumb]] | ||
[[File:Sanner96.pdf]] | [[File:Sanner96.pdf| thumb]] | ||
[[File:Sanner2.pdf]] | [[File:Sanner2.pdf| thumb]] | ||
[[File:Bajaj1999.pdf]] | [[File:Bajaj1999.pdf| thumb]] | ||
[[File:jp2007.pdf]] | [[File:jp2007.pdf| thumb]] | ||
[[File:Cossi96.pdf]] | [[File:Cossi96.pdf| thumb]] | ||
= References = | = References = | ||
Latest revision as of 15:55, 10 May 2022
Introduction
Calculating molecular surfaces is important and useful for structure-based drug design, representing molecules and computing inter-molecular interactions. It is also useful for displaying certain molecular properties on the surface, using color-coded and texture maps. Examples of such properties include atomic charge, electrostatic potential, hydrophobicity, polarizability, etc.
Types of Molecular Surfaces
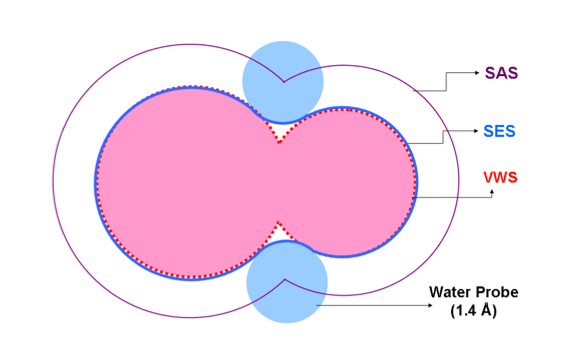
Van der Waals Surface
This surface is a union-of-balls representation. While acceptable for small proteins such as ligands, it fails to provide a good enough representation for larger proteins, returning too much buried surface.
Solvent Accessible Surface (SAS)
The SAS is also know as the Lee-Richards surface [1], and is calculated using the "rolling ball" algorithm developed by Shrake and Rupley [2], in which a sphere representing the solvent is used to "probe" the surface of the molecule. An important consideration is the selection of the probe radius: if it is too large, the resulting surface tends to be smaller and less accurate, whereas a smaller radius gives a larger surface with more details (bumps, cavities, etc.). A radius of 1.4 Å is normally chosen, approximating water. Also important are the assignment of van der Waals radii, the quality of the atomic coordinates, and hydrogen atom positions.
Solvent Excluded Surface (SES)
An improvement over the Lee-Richards SAS surface, Richards' SES [3] was implemented by Connolly in 1983 and so, is also known as the Connolly surface[4]. The following two parts are computed:
- contact surface, which is part of the van der Waals surface
- reentrant surface, which is the inward facing surface of the probe when it is in contact with more than one surface atom
Analytical Molecular Surface Calculation Algorithm
This algorithm for calculating solvent excluded surfaces was presented by Connolly in 1983 [5].
Surface creation
The rolling of a probe sphere is best described in terms of translational degrees of freedom: for each atom the probe touches, it loses one degree of freedom. The main idea in the algorithm is to start with probe positions with the degree of freedom equal to zero, and then to work one's way up; lower degrees of freedom provide starting and stopping points for higher degrees.
Accordingly, there are three different types of surface pieces.
For each case, a different piece of surface is generated, defined by the following:
- sphere: center, radius
- torus: center, 2 radii, axial vector
- boundary (circular arcs): center, radius, plane, end points
All of these pieces form a connected network, like a polyhedron where each piece is a face. The join between these faces is smooth, i.e. there is a well-defined tangent plane at all the points on an edge. This is in contrast with the van der Waals surface, where there are singularities.
The equations defining the surface are:
Torus construction
In order to build concave and saddle faces, a torus must be calculated for every pair of neighboring atoms. Two atoms i and j are neighbors if: .
When the probe is rolled over i and j, its center traces a circle and its surface traces the volume of a torus. The surface now connecting the two atoms is saddle-shaped.
The torus is described by its:
- center: circle center
- radius: circle radius
- plane: plane circle lies in
Probe placement and concave faces
Along the torus central circle, there may be other atoms k, such that k is a mutual neighbor of i and j. In this case there are two possible probe placements, one on each side of the plane passing through the three atom centers. If the third atom k is too far away for the probe to collide with it, however, or if k is located between i and j such that the probe has a van der Waals clash with it, no probe placement is possible.
Saddle faces
Once concave edges have been generated, tori are examined to create saddle faces. Three cases occur:
- buried tori generate no surface
- free tori (special case: no concave edges, and convex edges form full contact circle)
- partially accessible tori: one or more saddle-shaped rectangles are formed by choosing concave arcs that are adjacent in space and separated by convex arcs.
Convex faces
There are three types of atoms:
- buried, for which no surface is generated
- entirely accessible, which have no neighbors and a null edge list
- partially accessible, which have an edge list of size 1 or more
For a given atom, convex edges belonging to its edge list form a cycle, which has an orientation (counter-clockwise). The interior of the cycle is part of the van der Waals surface of the atom (left of edge).
Self-intersecting surfaces
Such intersections, also known as singularities, which cause surfaces to become discontinued or non-smooth, may occur in the following two conditions:
- a saddle face intersects itself if
- two concave faces interpenetrate
These problems usually happen in deep grooves. Sanner provides a method for dealing with such problems in MSMS.
Fast and Robust Computation of Molecular Surfaces
In MSMS, Sanner extends previous work by Connolly by introducing the r-reduced surface, and presenting a method for handling singularities [6] [7].
The r-reduced surface
The r-reduced surface is equivalent to the -shape, where . Both the SAS and SES can be recovered from the r-reduced surface.
This surface is defined by the following:
- face: polygon obtained by connecting centers of atoms probe is lying on
- vertices: atom centers
- edges: lines connecting vertices
An example reduced surface looks like this:
To each face corresponds a spheric reentrant face of SES, and for each edge, a toric reentrant face. Cycles of convex edges define contact faces.
Examples of molecular surface images: http://www.scripps.edu/~sanner/html/ms_gallery.html
Handling singularities
- Radial singularities are detected during computation of r-reduced surface, and stored with edge. A toric reentrant face is then made out of the 2 triangular faces connected by a singular vertex or edge.
- Non-radial singularities are handled in three step:
- spheric reentrant faces are clipped
- if a singular edge is inside a probe, it is entirely or partially "eaten"; singular vertices are added at the intersection of the probes, and the eaten part of the edge is replaced with singular edges lying on the intersection circles on the probes
- for each pair of tangent probes generating spheric reentrant faces, not tangent to the same 3 atoms, clip faces at intersection circle C
Multiresolution Molecular Shapes
This algorithm was presented by Bajaj et al. in 1999 [8].
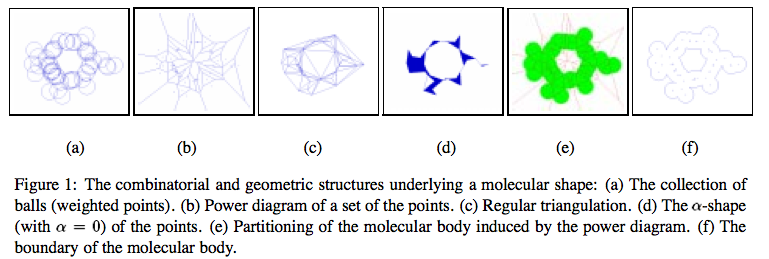
Supporting geometric structures
- Power diagram: tiling of space into convex regions where tile i is the set of points nearest to vertex Pi, in power distance metric.
- Regular triangulation: dual of the power diagram, like the Delauney triangulation is the dual of the Voronoi diagram.
- Weighted alpha shapes: a simplex s in the regular triangulation belongs to the alpha shape if the weighted point orthogonal to the vertices of s is smaller than alpha. Example: an edge (u,v) is part of the zero-shape if the two balls centered at u and v intersect.
Decimation of molecular shapes and construction of multiresolution hierarchy
The goal of decimating a molecular structure is to produce a coarser and simpler representation of the model in order to reduce computational costs. This operation is performed on the set of weighted points, which makes it easier to have a hierarchical representation for multiple types of molecular shapes.
Coarsening is transitive. This allows for the construction of an efficient multiresolution representation of molecular models.
Vertex clustering is performed: if two balls overlap (i.e. the edge joining their centers belong to the zero-shape), then they are removed from the triangulation, and a new ball is inserted instead.
Two different clustering techniques can be used:
- Local Vertex Clustering (LVC) (left)
- Minimal Vertex Clustering (MVC) (right)
LVC is simpler and more efficient, but MVC returns a tighter coarsening.
The algorithm for building the multiresolution hierarchy depends on this clustering of edges in the zero-shape, creating consecutive levels of coarseness.
Examples
Application to Implicit Nonpolar Solvent Models
Implicit solvation models generally have two components:
- electrostatic, or polar, which can be calculated using Poisson-Boltzmann model
- non-electrostatic, or nonpolar, which can be estimated using SAS:
This model is widely used, although its correlation with the explicit solvent model is rather poor. It could be improved upon [9] by breaking down the nonpolar energy term into: and writing each one as follows:
is the solvent-solute distance and is the van der Waals attractive potential.
Software Packages
- Molecular Surface Package (MSP), M.L. Connolly 1983
- SURFNET, R.A. Laskowski 1995
- Molecular Shape Generation, C.L. Bajaj 1999
- The Analytic Surface Calculation Package (ASC), F. Eisenhaber 1993
Example Using MSMS
Download
- Binary versions of MSMS are available for most platforms at: http://mgltools.scripps.edu/downloads#msms
- Included in the distribution are the following executables:
- msms."arch": Binary version of MSMS compiled for "arch"
- pdb_to_xyzr: Script to convert get XYZR coordinates from a PDB file
- pdb_to_xyzrn: Script to convert get XYZRN coordinates from a PDB file
Input
- MSMS takes XYZR (or XYZRN) files as input. The main difference between the two types of input files is that XYZRN files also contain the names of each atom at the end of the line, in addition to atomic coordinates and van der Waals radii. This can be useful if you want the surface on a per-atom basis.
- Input files can be obtained by executing the following commands on PDB files:
$ pdb_to_xyzr 1HSG_r.pdb > 1HSG_r.xyzr
1HSG_r.xyzr 29.361 39.686 5.862 1.70 30.307 38.663 5.319 2.00 29.760 38.071 4.022 1.74 28.600 38.302 3.676 1.40 30.508 37.541 6.342 2.00 29.296 37.591 7.162 2.00 28.778 39.015 7.019 2.00 ...
$ pdb_to_xyzrn 1HSG_r.pdb > 1HSG_r.xyzrn
1HSG_r.xyzrn 29.361000 39.686000 5.862000 1.700000 1 N_PRO_1 30.307000 38.663000 5.319000 2.000000 1 CA_PRO_1 29.760000 38.071000 4.022000 1.740000 1 C_PRO_1 28.600000 38.302000 3.676000 1.400000 1 O_PRO_1 30.508000 37.541000 6.342000 2.000000 1 CB_PRO_1 29.296000 37.591000 7.162000 2.000000 1 CG_PRO_1 28.778000 39.015000 7.019000 2.000000 1 CD_PRO_1 ...
Generating the surface
- A man page is available online: http://www.scripps.edu/~sanner/html/msms_man.html
- After generating the input file, MSMS is run by:
$ msms -if 1HSG_r.xyzr -of 1HSG_r
This generates two files: 1HSG_r.face (triangulated surface faces) and 1HSG_r.vert (vertices).
- To obtain the surface on a per-atom basis, run:
$ msms -if 1HSG_r.xyzrn -of 1HSG_r -af 1HSG_r
- Using the -af option, an additional file is created: 1HSG_r.area, which contains the surface area on a per-atom basis.
$ msms.x86_64Linux2.2.6.1.staticgcc -if 1HSG_r.xyzr -of 1HSG_r
MSMS 2.6.1
Copyright M.F. Sanner (1994)
Compilation flags -O2 -DVERBOSE -DTIMING
INPUT 1HSG_r.xyzr 1514 spheres 0 collision only, radii 1.400 to 2.000
PARAM Probe_radius 1.500 density 1.000 hdensity 3.000
REDUCED SURFACE ...
RS component 0 identified 343 344 341
0 free edge(s) found in component 0
RS component #faces #edges #free_edges #vertices genus
0 1738 2607 0 817 27
Time Reduced Surface real: 0.04 user: 0.05 sys: 0.00
ANALYTICAL SOLVENT EXCLUDED SURFACE...
Component 0
Time Surface real: 0.01 user: 0.01 sys: 0.00
Time Singularities real: 0.00 user: 0.00 sys: 0.00
SES comp. #fac. #edg. #s_edg. #vert. #s_vert. R_h C_h genus
0 5314 10716 107 5402 188 0 0 1
ANALYTICAL SURFACE AREA :
Comp. probe_radius, reent, toric, contact SES SAS
0 1.500 2190.425 3376.937 2929.180 8496.541 9767.748
TRIANGULATION...
Component 0
component#, vertices, edges, faces, genus, density
0 9600 28800 19200 1 1.130
Time Triangulation real: 0.03 user: 0.04 sys: 0.00
NUMERICAL VOLUMES AND AREA
Comp. probe_radius SES_volume SES_area)
0 1.50 26064.947 8129.946
Total ses_volume: 26064.947
MSMS terminated normally
Total Time real: 0.16 user: 0.14 sys: 0.01
Visualizing the output
MSMS output can be visualized using a number of graphics tools.
This example is displayed using DINO 3D (http://www.dino3d.org/intro.php).
// load the structure dataset load 1HSG_r.pdb -name 1HSG_r // load the surface dataset load 1HSG_r -type msms -name surf // attach the surface to the structure // (thereby assigning each surface point // to an atom) .surf attach .1HSG_r // center on the surface scene center [.surf] // create a surface object .surf new -name all // adjust the clipping planes scene autoslab

// color the surface object according // to some underlying residue properties // the variables used here are predefined // by DINO, use 'var' to look at them .surf.all set color=blue -sel $basic .surf.all set color=blue4 -sel $basic2 .surf.all set color=red -sel $acidic .surf.all set color=red4 -sel $acidic2 .surf.all set color=cyan -sel $polar .surf.all set color=green -sel $aromatic .surf.all set color=yellow -sel $aliphatic

// reset the color to white and make the surface // transparent to show the underlying structure .surf.all set color=white .1HSG_r new -name all .1HSG_r.all render custom .surf.all render t=0.4
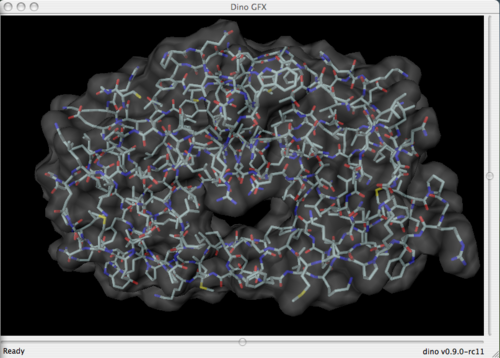
More tutorials are available at: http://www.dino3d.org/tutorials/
Further Reading








References
- ↑ Lee, B. and Richards, F.M., The Interpretation of Protein Structures: Estimation of Static Accessibility, 1971. J. Appl. Cryst. 55: p. 379-400.
- ↑ Shrake A. and Rupley J.A., Environment and exposure to solvent of protein atoms. Lysozyme and insulin, 1973. J. Mol. Biol. 79(2): p. 351-371.
- ↑ Richards, F.M., Areas, Volumes, Packing, and Protein Structure, 1977. Ann. Rev. Biophys. Bioeng. 6: p. 151-176.
- ↑ Connolly, M.L., Analytical Molecular Surface Calculation, 1983. J. Appl. Cryst. 16: p. 548-558.
- ↑ Connolly M.L., Analytical Molecular Surface Calculation, 1983. J. Appl. Cryst. 16: p. 548-558.
- ↑ Sanner, M., Olson, A.J. and Sephner, J.-C., Fast and Robust Computation of Molecular Surfaces, 1995. Proc. 11th ACM Symp. Comp. Geom.. C6-C7.
- ↑ Sanner, M., Olson, A.J. and Sephner, J.-C., Reduced Surface: An Efficient Way to Compute Molecular Surfaces, 1996. Biopolymers. 38. pp. 305-320.
- ↑ Bajaj, C.L., Pascucci, V. Shamir, A., Holt, R.J. and Netravali A.N., Multiresolution Molecular Shapes, 1999. TICAM Report. 99-42.
- ↑ Tan, C., Tan Y.H. and Luo R., Implicit Nonpolar Solvent Models, 2007. J. Phys. Chem. B. 111. pp. 12263-12274.